Welcome to Viking’s Documentation¶
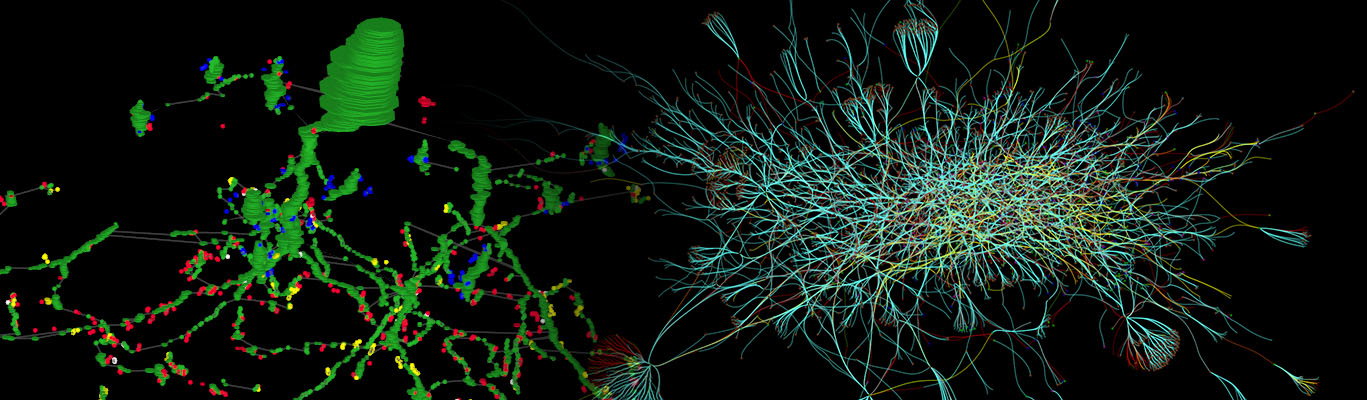
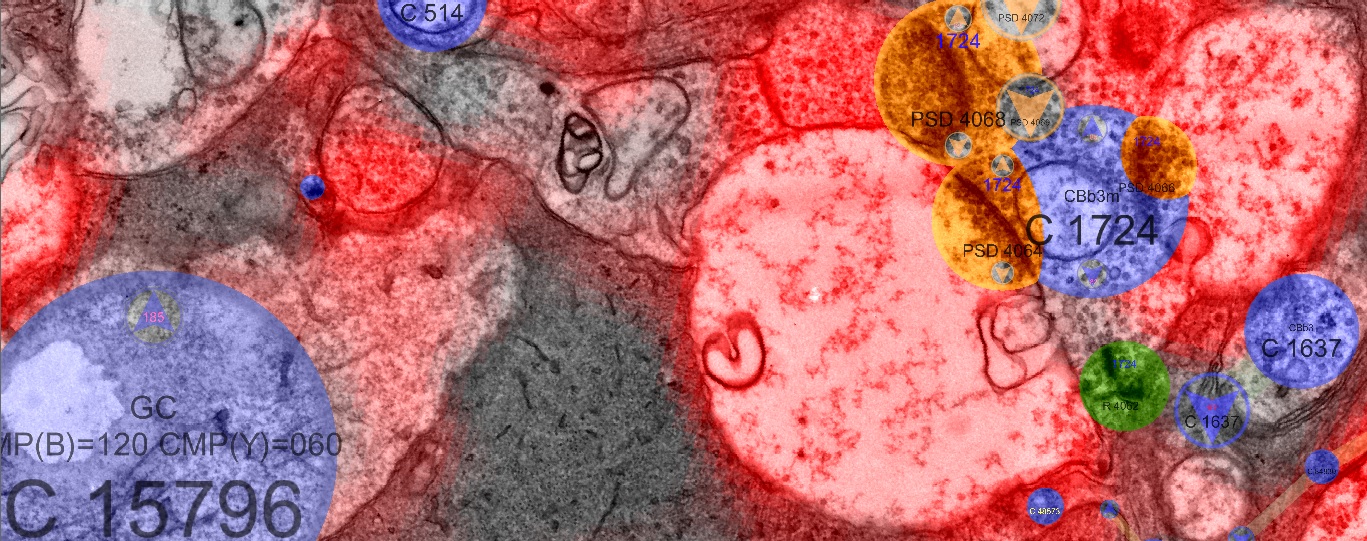
Viking is a multi-user web-based collaborative management system for images and volumes which allows users to view multi-terabyte datasets, annotate images with their own annotation schema, and summarize the results. Viking was developed by James Anderson at the Marc Lab for use with the first retinal connectome which was assembled using SSTEM and Computation Molecular Phenotyping
Viking’s role begins after images are collected and registered into a volume. The Marc Lab uses Nornir to build our volumes.
Viking Client Installation¶
- Viking requires the following:
-
-
Windows 7+
-
.NET Framework 4.8
-
A dedicated GPU. Intel chips with embedded graphics also work.
-
An internet connection
-
Click here to install the Viking client ¶
Note for existing users: July 2023 - There was an issue with Chrome where it would not download the most recent version of setup.exe using the link above and would instead pull from a cache. If setup isn’t working try another browser or clear cached files from your browser history. Also, sometime in June 2023 the web installer for .NET 4.8.1 automatically downloaded from Microsoft is no longer valid. You will need to manually install .NET 4.8.1 if it is not on your system, but a current Windows install should include it.
Note for IT Administrators: Viking’s installer is signed by the University of Utah. Adding Viking’s public key as a trusted publisher allows users to install Viking without administrator rights.
Contents¶
Account creation¶
Accounts are only required to write to the database. Read-only access via Viking may be done anonymously.
Create accounts and do some basic visualization on the original visualization site